Report the individuals among turtles that are on the same patches as
the agents.
turtlesOn(world, turtles, agents, breed, simplify = TRUE)
# S4 method for worldNLR,agentMatrix,matrix,missing
turtlesOn(world, turtles, agents, simplify)
# S4 method for worldNLR,agentMatrix,matrix,character
turtlesOn(world, turtles, agents, breed, simplify = TRUE)Arguments
- world
WorldMatrixorworldArrayobject.- turtles
AgentMatrixobject representing the movingagents.- agents
Matrix (
ncol= 2) with the first columnpxcorand the second columnpycorrepresenting thepatchescoordinates, or- breed
Characters. Vector of
breednames for the selectedturtles. If missing, there is no distinction based uponbreed.- simplify
Logical. If
simplify = TRUE, allturtleson the samepatchesas anyagentsare returned; ifsimplify = FALSE, theturtlesare evaluated for eachagents'spatchesindividually.
Value
AgentMatrix representing any individuals from turtles of
any of the given breed, if specified,
located on the same patches as any of the agents, if simplify = TRUE, or
Matrix (`ncol` = 2) with the first column `whoTurtles` and the second column
`id` showing which `turtles` are on the same
`patches` as which `agents` represented by `id`, if `simplify = FALSE`.
`id` represents and follows the order of the `agents`. `id` does not represent
the `who` numbers
of the `agents` if `agents` are `turtles`.Details
The agents must be located inside the
world's extent.
References
Wilensky, U. 1999. NetLogo. http://ccl.northwestern.edu/netlogo/. Center for Connected Learning and Computer-Based Modeling, Northwestern University. Evanston, IL.
Examples
w1 <- createWorld(
minPxcor = 0, maxPxcor = 9, minPycor = 0, maxPycor = 9,
data = runif(100)
)
t1 <- createTurtles(n = 500, coords = randomXYcor(w1, n = 500))
plot(w1)
points(t1, col = of(agents = t1, var = "color"), pch = 16)
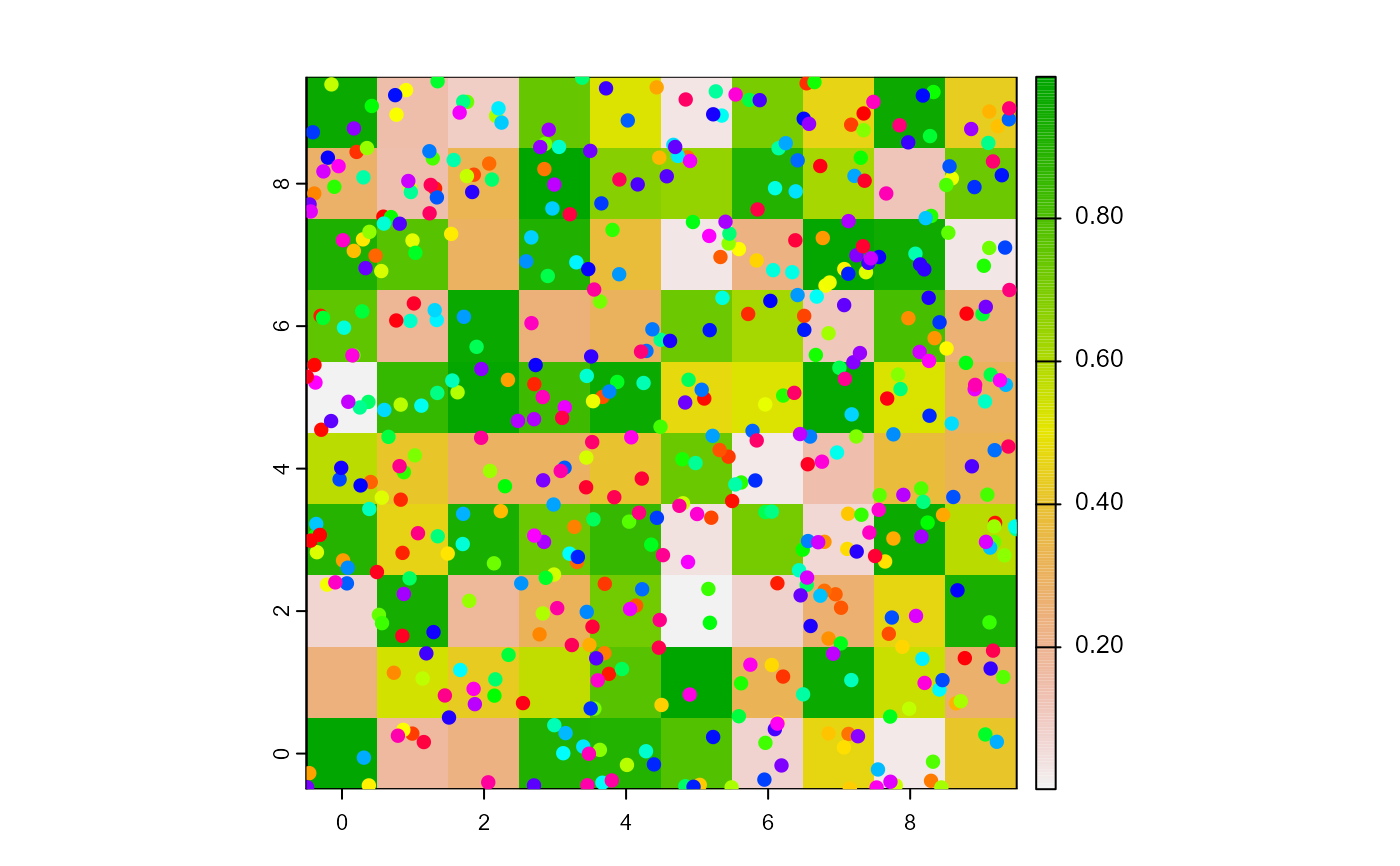 t2 <- turtlesOn(world = w1, turtles = t1, agents = patch(w1, 2, 2))
t2 <- turtlesOn(world = w1, turtles = t1, agents = patch(w1, 2, 2))